seqspec, short for “sequence specification” (pronounced “seek-speck”), is a file format that describes data generated from genomics experiments. Both the file format and seqspec tool enable uniform processing of genomics data.
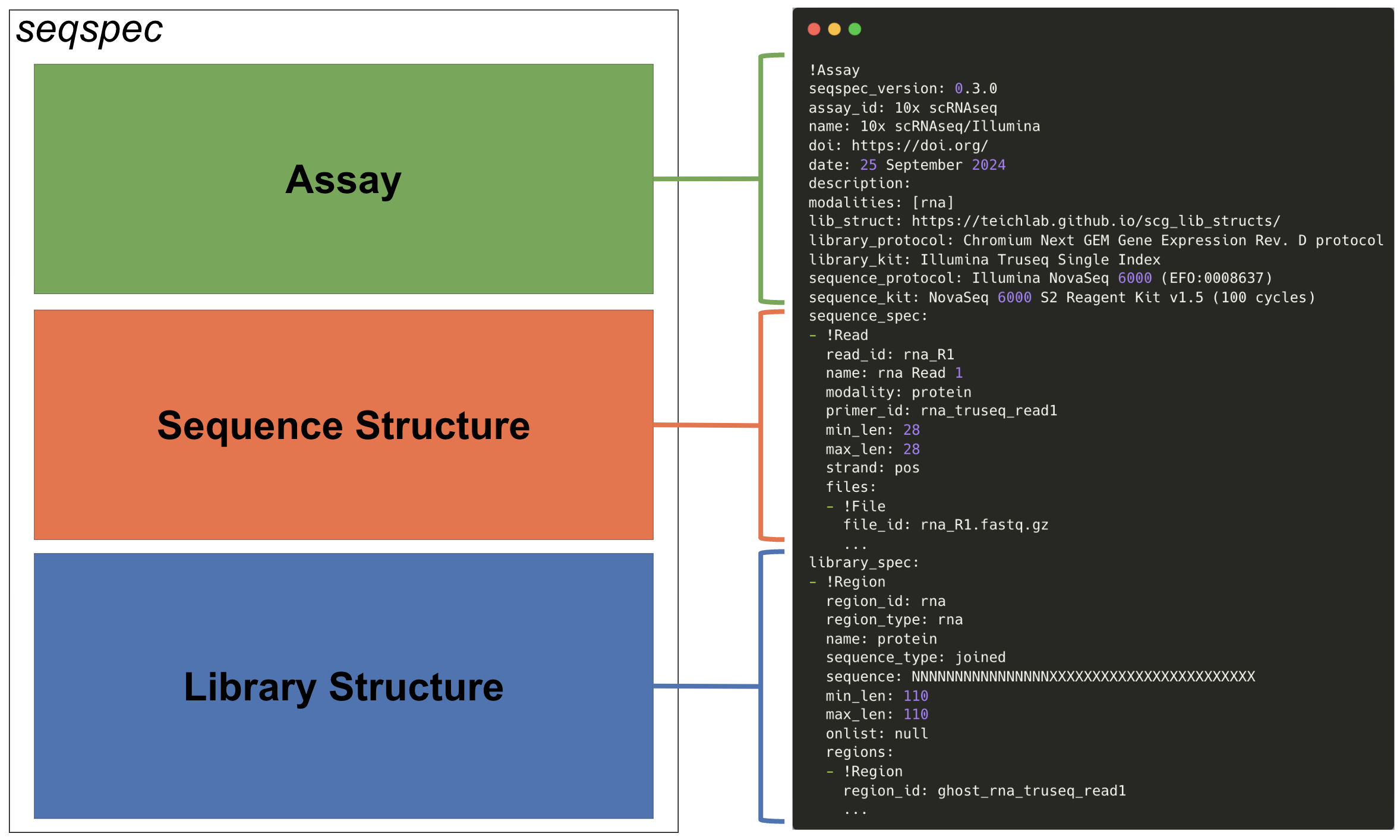 Figure 1: Anatomy of a
Figure 1: Anatomy of a seqspec file.
We have multiple tutorials to get you up and running with seqspec:
Learn how to use
seqspecto standardize your genomics data preprocessing.Understand how to manipulate
seqspecfiles using theseqspeccommand-line tool.
Current release¶
seqspec 0.4.0 keeps the Python and Rust implementations aligned around the same core command set.
seqspec upgradeupgrades0.3.0specs to0.4.0in both implementations.seqspecloads gzipped specs directly, so.yaml.gzworks anywhere a spec path is accepted.seqspec authmanages host-matched auth profiles for remote resources, andseqspec check/seqspec onlistcan use them with--auth-profile.seqspec checknow emits warnings for valid but risky geometry, such as reads that cover the same declared regions.seqspec onlist -s region-typenow errors when the same region type appears across multiple reads, so ambiguous joins are explicit.seqspec print -f seqspec-htmlwrites a self-contained HTML view of the library and reads.seqspec buildis deprecated.
Citation¶
The seqspec format and tool are described in this publication. If you use seqspec please cite
Ali Sina Booeshaghi, Xi Chen, Lior Pachter, A machine-readable specification for genomics assays, Bioinformatics, Volume 40, Issue 4, April 2024, btae168.seqspec was inspired by and builds off of the Teichmann Lab Single Cell Genomics Library Structure by Xi Chen.
Documentation¶
Learn about the
seqspecspecification:docs/SPECIFICATION.mdWrite a
seqspecfrom a simple example:docs/TUTORIAL_SIMPLE.mdWrite a
seqspecfrom a template:docs/TUTORIAL_FROM_TEMPLATE.mdBrowse the generated example site:
docs/examples/site/assays.html
Rust implementation¶
auth : Manage remote authentication profiles.
build : Deprecated in both CLIs.
check : Validate seqspec file against specification (verify check)
find : Find objects in seqspec file
file : List files present in seqspec file
format : Autoformat seqspec file
index : Identify position of elements in seqspec file
info : Get information from seqspec file
init : Generate a new empty seqspec file
insert : Insert regions or reads into an existing spec (TODO: move Input structs to models)
methods : Convert seqspec file into methods section
modify : Modify attributes of various elements in seqspec file
onlist : Get onlist file for elements in seqspec file
print : Display the sequence and/or library structure from seqspec file
split : Split seqspec file by modality
upgrade : Upgrade seqspec file to current version
version: Get seqspec tool version and seqspec file version
The standalone Rust CLI supports library-ascii, seqspec-ascii, and seqspec-html in seqspec print. seqspec-png remains Python-only for now.
- Booeshaghi, A. S., Chen, X., & Pachter, L. (2024). A machine-readable specification for genomics assays. Bioinformatics, 40(4). 10.1093/bioinformatics/btae168
- Booeshaghi, A. S., Chen, X., & Pachter, L. (2024). A machine-readable specification for genomics assays. Bioinformatics, 40(4). 10.1093/bioinformatics/btae168